Cluster no. 53
Go back to cluster table
Cluster is part of supercluster: 1
Cluster characteristics:
| size |
20029 |
| size_real |
20029 |
| ecount |
11396680 |
| supercluster |
1 |
| annotations_summary |
|
| pair_completeness |
0.563544106167057 |
| pbs_score |
None |
| TR_score |
None |
| TR_monomer_length |
None |
| loop_index |
0.0844775076139598 |
| satellite_probability |
6.63894134171845e-17 |
| consensus |
None |
| TAREAN_annotation |
Other |
| orientation_score |
1 |
protein domains:
No protein domains detected
protein domains:

Reads annotation summary
No similarity hits to repeat databases found
clusters with similarity:
| Cluster |
Number of similarity hits |
|
| 55 |
388000 |
| 79 |
68300 |
| 43 |
7600 |
| 190 |
1050 |
| 102 |
119 |
| 117 |
37 |
| 35 |
1 |
| 52 |
1 |
| 100 |
1 |
|
clusters connected through mates:
| Cluster |
Number of shared
read pairs |
k |
|
| 55 |
2280 |
0.355 |
| 79 |
2180 |
0.425 |
| 43 |
361 |
0.0679 |
| 35 |
69 |
0.0116 |
| 31 |
15 |
0.00195 |
| 41 |
11 |
0.00194 |
| 4 |
9 |
0.00147 |
| 102 |
9 |
0.00177 |
| 117 |
9 |
0.00228 |
| 6 |
8 |
0.00123 |
| 190 |
8 |
0.00226 |
| 1 |
7 |
0.00113 |
| 29 |
7 |
0.00154 |
| 52 |
7 |
0.00153 |
| 108 |
7 |
0.00196 |
| 17 |
6 |
0.000711 |
| 19 |
6 |
0.0013 |
| 27 |
6 |
0.00147 |
| 38 |
6 |
0.00152 |
| 39 |
6 |
0.00143 |
| 67 |
6 |
0.00123 |
| 15 |
5 |
0.000982 |
| 26 |
5 |
0.00104 |
| 36 |
5 |
0.00117 |
| 58 |
5 |
0.00139 |
| 66 |
5 |
0.000904 |
| 72 |
5 |
0.00124 |
| 77 |
5 |
0.00124 |
| 3 |
4 |
0.000632 |
| 10 |
4 |
0.00097 |
| 11 |
4 |
0.000641 |
| 25 |
4 |
0.000876 |
| 33 |
4 |
0.000984 |
| 48 |
4 |
0.000673 |
| 74 |
4 |
0.000951 |
| 119 |
4 |
0.0012 |
| 182 |
4 |
0.00112 |
| 2 |
3 |
0.000364 |
| 5 |
3 |
0.000695 |
| 16 |
3 |
0.000571 |
| 30 |
3 |
0.000475 |
| 45 |
3 |
0.000558 |
| 46 |
3 |
0.000695 |
| 49 |
3 |
0.000677 |
| 51 |
3 |
0.000537 |
| 92 |
3 |
0.000885 |
| 100 |
3 |
0.000809 |
| 105 |
3 |
0.000713 |
| 111 |
3 |
0.000568 |
| 112 |
3 |
0.000872 |
| 126 |
3 |
0.000769 |
| 130 |
3 |
0.000626 |
| 135 |
3 |
0.000803 |
| 138 |
3 |
0.000855 |
| 153 |
3 |
0.000738 |
| 169 |
3 |
0.000855 |
| 180 |
3 |
0.000939 |
| 20 |
2 |
0.000326 |
| 23 |
2 |
0.000395 |
| 24 |
2 |
0.000347 |
| 28 |
2 |
0.000479 |
| 34 |
2 |
0.000393 |
| 44 |
2 |
0.00035 |
| 50 |
2 |
0.000457 |
| 59 |
2 |
0.000519 |
| 70 |
2 |
0.000551 |
| 94 |
2 |
0.000647 |
| 97 |
2 |
0.000463 |
| 98 |
2 |
0.000547 |
| 106 |
2 |
0.000481 |
| 109 |
2 |
0.000576 |
| 110 |
2 |
0.000549 |
| 113 |
2 |
0.000494 |
| 114 |
2 |
0.000586 |
| 121 |
2 |
0.000607 |
| 134 |
2 |
0.000498 |
| 164 |
2 |
0.000665 |
| 173 |
2 |
0.000611 |
| 187 |
2 |
0.000586 |
| 189 |
2 |
0.000535 |
| 213 |
2 |
0.000518 |
| 220 |
2 |
0.000629 |
| 238 |
2 |
0.00065 |
| 245 |
2 |
0.00059 |
| 359 |
2 |
0.000709 |
| 9160 |
2 |
0.000715 |
| 7 |
1 |
0.000139 |
| 9 |
1 |
0.000187 |
| 12 |
1 |
0.000148 |
| 14 |
1 |
0.000165 |
| 21 |
1 |
0.000198 |
| 22 |
1 |
0.000184 |
| 32 |
1 |
0.000237 |
| 37 |
1 |
0.000172 |
| 40 |
1 |
0.00016 |
| 42 |
1 |
0.00022 |
| 56 |
1 |
0.000331 |
| 57 |
1 |
0.000264 |
| 60 |
1 |
0.000261 |
| 61 |
1 |
0.000228 |
| 62 |
1 |
0.000185 |
| 65 |
1 |
0.000247 |
| 71 |
1 |
0.000201 |
| 75 |
1 |
0.000175 |
| 76 |
1 |
0.000244 |
| 80 |
1 |
0.000244 |
| 81 |
1 |
0.000201 |
| 82 |
1 |
0.000267 |
| 83 |
1 |
0.000253 |
| 84 |
1 |
0.000232 |
| 86 |
1 |
0.000193 |
| 89 |
1 |
0.000245 |
| 91 |
1 |
0.00029 |
| 99 |
1 |
0.00029 |
| 101 |
1 |
0.000234 |
| 103 |
1 |
0.000211 |
| 107 |
1 |
0.000287 |
| 115 |
1 |
0.000222 |
| 116 |
1 |
0.000208 |
| 118 |
1 |
0.00028 |
| 122 |
1 |
0.000251 |
| 127 |
1 |
0.000253 |
| 131 |
1 |
0.000328 |
| 132 |
1 |
0.000284 |
| 133 |
1 |
0.000182 |
| 139 |
1 |
0.000258 |
| 144 |
1 |
0.000304 |
| 145 |
1 |
0.000239 |
| 146 |
1 |
0.000285 |
| 155 |
1 |
0.000285 |
| 156 |
1 |
0.000277 |
| 157 |
1 |
0.000281 |
| 159 |
1 |
0.000279 |
| 161 |
1 |
0.000245 |
| 163 |
1 |
0.000247 |
| 166 |
1 |
0.000289 |
| 167 |
1 |
0.000244 |
| 168 |
1 |
0.000255 |
| 171 |
1 |
0.000276 |
| 172 |
1 |
0.000254 |
| 174 |
1 |
0.00026 |
| 181 |
1 |
0.000283 |
| 186 |
1 |
0.000247 |
| 188 |
1 |
0.000345 |
| 192 |
1 |
0.000344 |
| 193 |
1 |
0.000309 |
| 194 |
1 |
0.000336 |
| 195 |
1 |
0.000317 |
| 197 |
1 |
0.000271 |
| 198 |
1 |
0.000313 |
| 203 |
1 |
3e-04 |
| 205 |
1 |
0.000293 |
| 211 |
1 |
0.000303 |
| 214 |
1 |
0.000322 |
| 216 |
1 |
0.000326 |
| 219 |
1 |
0.000323 |
| 222 |
1 |
0.000317 |
| 223 |
1 |
0.00032 |
| 224 |
1 |
0.000347 |
| 230 |
1 |
0.000339 |
| 236 |
1 |
0.000351 |
| 246 |
1 |
0.000348 |
| 251 |
1 |
0.000337 |
| 255 |
1 |
0.000309 |
| 262 |
1 |
0.000351 |
| 269 |
1 |
0.000342 |
| 282 |
1 |
0.000339 |
| 293 |
1 |
0.000342 |
| 300 |
1 |
0.000346 |
| 305 |
1 |
0.000352 |
| 308 |
1 |
0.000347 |
| 309 |
1 |
0.000352 |
| 390 |
1 |
0.000352 |
| 398 |
1 |
0.000355 |
| 418 |
1 |
0.00035 |
| 817 |
1 |
0.000355 |
| 1450 |
1 |
0.000356 |
| 1880 |
1 |
0.000356 |
| 2010 |
1 |
0.000356 |
| 2130 |
1 |
0.000357 |
| 2260 |
1 |
0.000357 |
| 2760 |
1 |
0.000357 |
| 3140 |
1 |
0.000357 |
| 3170 |
1 |
0.000357 |
| 4100 |
1 |
0.000357 |
| 4890 |
1 |
0.000357 |
| 7920 |
1 |
0.000357 |
| 8280 |
1 |
0.000357 |
| 14000 |
1 |
0.000357 |
| 15000 |
1 |
0.000358 |
| 16800 |
1 |
0.000357 |
| 16900 |
1 |
0.000357 |
| 17300 |
1 |
0.000357 |
| 21200 |
1 |
0.000358 |
| 22600 |
1 |
0.000358 |
| 24900 |
1 |
0.000358 |
| 25200 |
1 |
0.000358 |
| 26000 |
1 |
0.000358 |
| 37300 |
1 |
0.000357 |
| 39600 |
1 |
0.000357 |
| 43000 |
1 |
0.000358 |
| 44500 |
1 |
0.000357 |
| 45400 |
1 |
0.000358 |
| 48200 |
1 |
0.000358 |
| 55000 |
1 |
0.000358 |
| 57700 |
1 |
0.000358 |
| 63600 |
1 |
0.000358 |
| 74700 |
1 |
0.000358 |
| 77100 |
1 |
0.000358 |
| 83900 |
1 |
0.000358 |
| 87300 |
1 |
0.000358 |
| 103000 |
1 |
0.000358 |
| 105000 |
1 |
0.000358 |
| 131000 |
1 |
0.000358 |
| 141000 |
1 |
0.000358 |
| 148000 |
1 |
0.000358 |
| 163000 |
1 |
0.000358 |
| 173000 |
1 |
0.000358 |
| 186000 |
1 |
0.000358 |
| 204000 |
1 |
0.000358 |
| 217000 |
1 |
0.000358 |
| 217000 |
1 |
0.000358 |
| 240000 |
1 |
0.000358 |
| 242000 |
1 |
0.000358 |
| 243000 |
1 |
0.000358 |
| 257000 |
1 |
0.000358 |
| 271000 |
1 |
0.000358 |
| 273000 |
1 |
0.000358 |
| 275000 |
1 |
0.000358 |
| 296000 |
1 |
0.000358 |
| 321000 |
1 |
0.000358 |
| 321000 |
1 |
0.000358 |
| 344000 |
1 |
0.000358 |
|
CL53 ----> CL55
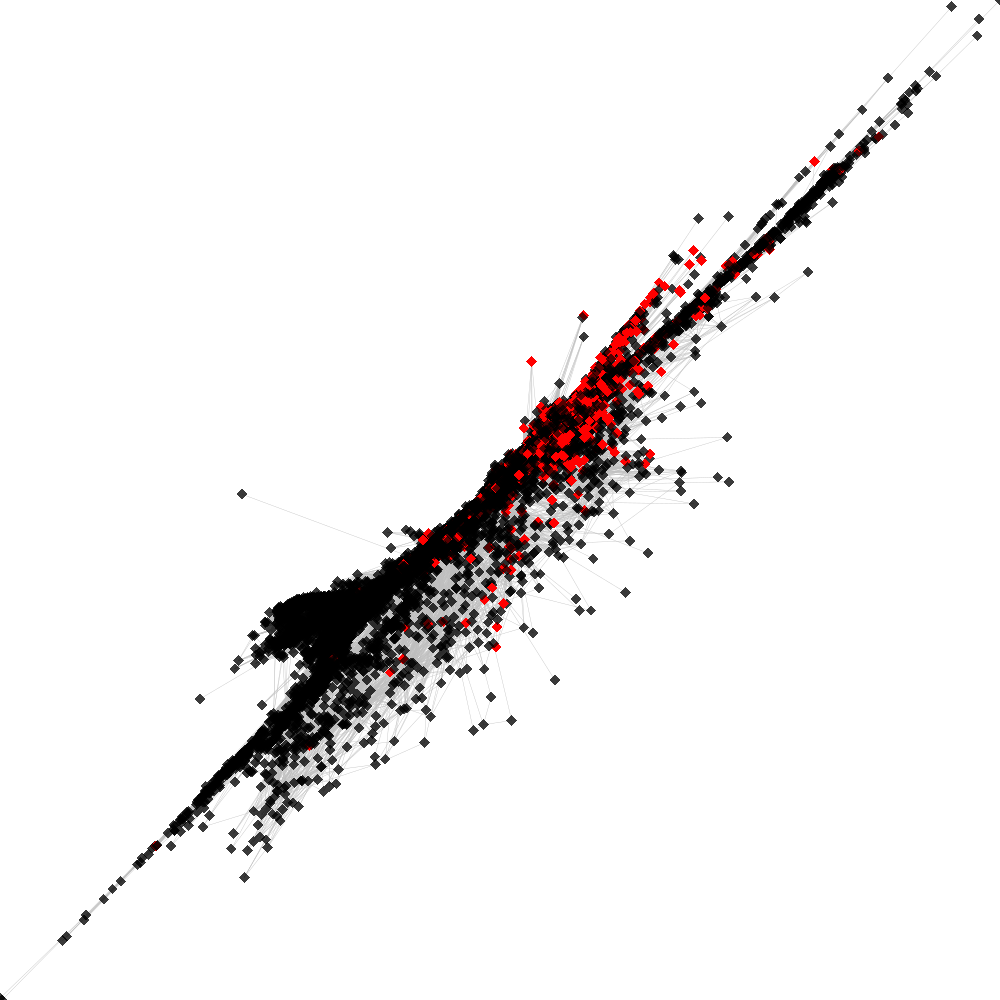
No. of shared pairs: :2282
CL53 ----> CL79
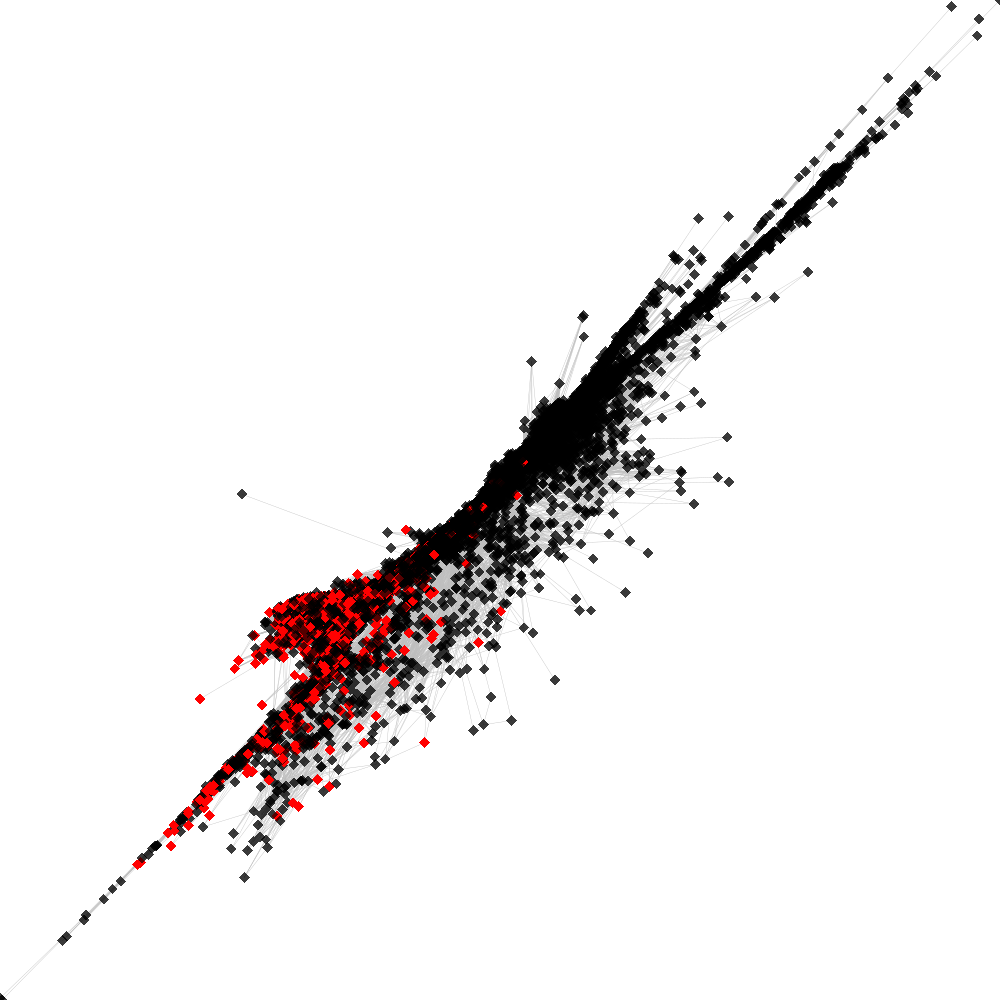
No. of shared pairs: :2182
CL53 ----> CL43
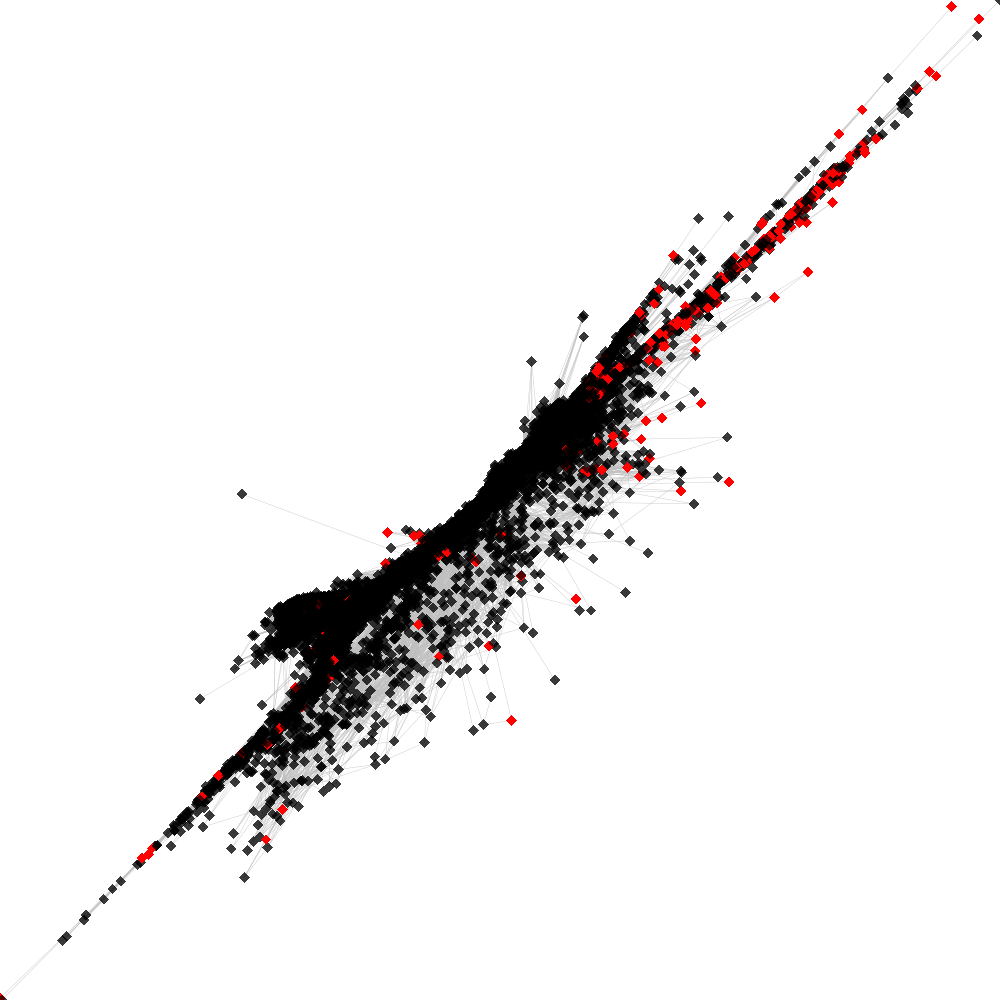
No. of shared pairs: :361
CL53 ----> CL35
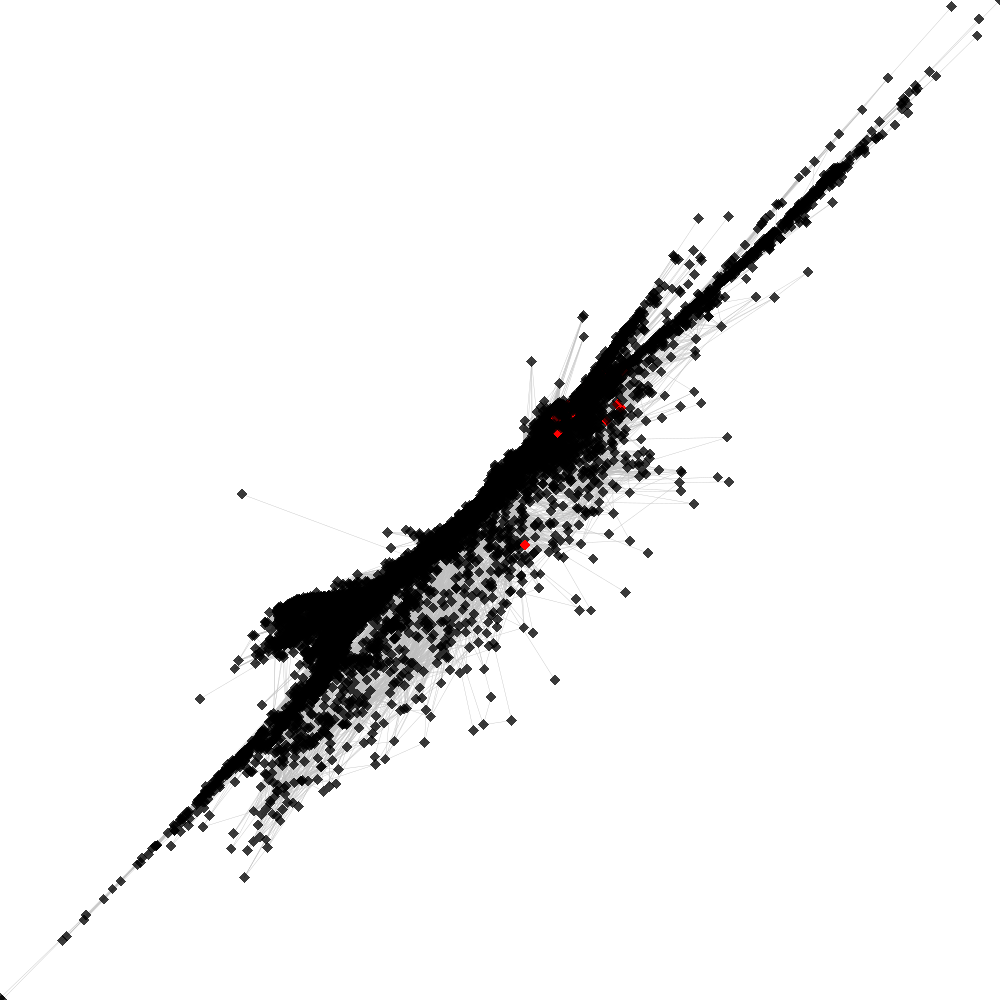
No. of shared pairs: :69
CL53 ----> CL31
No. of shared pairs: :15
CL53 ----> CL41
No. of shared pairs: :11

