Cluster no. 55
Go back to cluster table
Cluster is part of supercluster: 1
Cluster characteristics:
| size |
19433 |
| size_real |
19433 |
| ecount |
9646412 |
| supercluster |
1 |
| annotations_summary |
|
| pair_completeness |
0.454783650247043 |
| pbs_score |
None |
| TR_score |
None |
| TR_monomer_length |
None |
| loop_index |
0.0949415941954408 |
| satellite_probability |
6.46126039616052e-16 |
| consensus |
None |
| TAREAN_annotation |
Other |
| orientation_score |
1 |
protein domains:
No protein domains detected
protein domains:
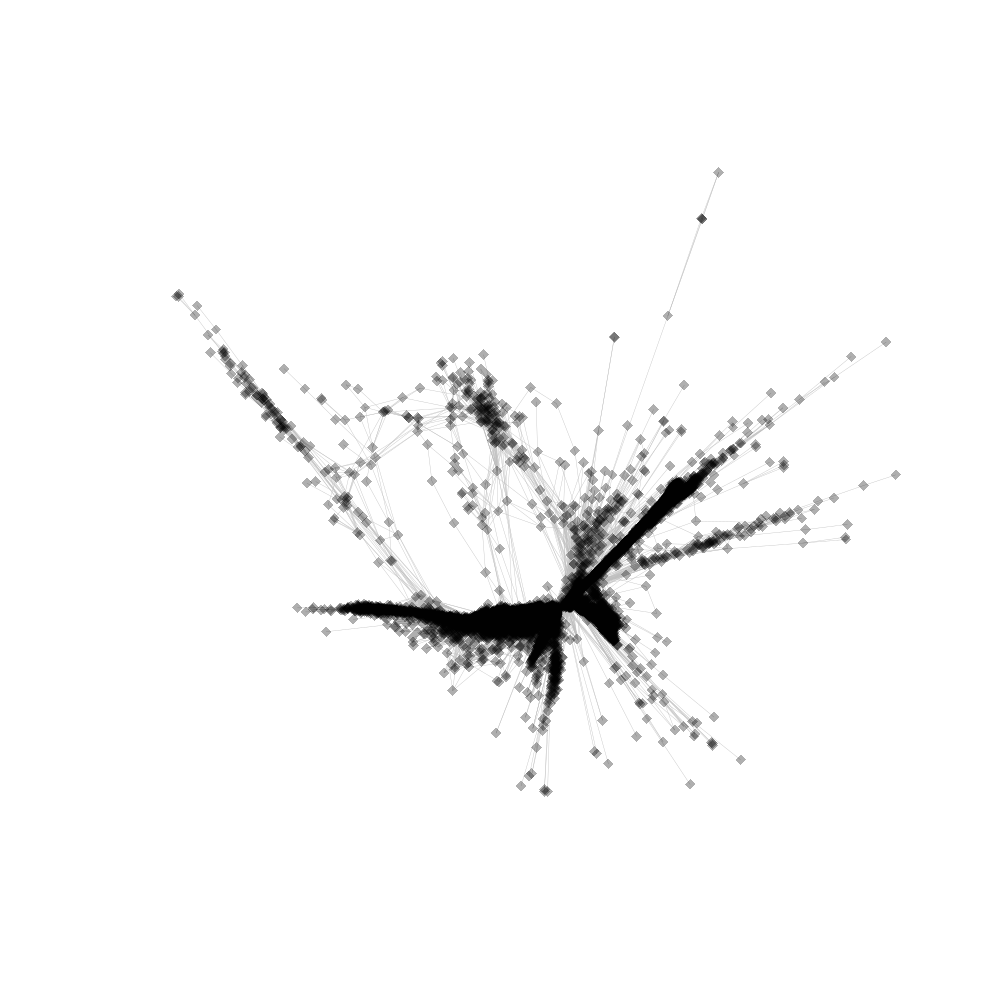

Reads annotation summary
No similarity hits to repeat databases found
clusters with similarity:
| Cluster |
Number of similarity hits |
|
| 53 |
388000 |
| 35 |
169000 |
| 190 |
28200 |
| 67 |
23200 |
| 117 |
4620 |
| 43 |
39 |
| 524 |
11 |
| 40 |
1 |
| 238 |
1 |
| 8100 |
1 |
|
clusters connected through mates:
| Cluster |
Number of shared
read pairs |
k |
|
| 35 |
2540 |
0.374 |
| 53 |
2280 |
0.355 |
| 67 |
878 |
0.153 |
| 190 |
619 |
0.141 |
| 43 |
123 |
0.02 |
| 117 |
72 |
0.015 |
| 524 |
22 |
0.00597 |
| 31 |
14 |
0.00164 |
| 4 |
10 |
0.00143 |
| 102 |
10 |
0.00169 |
| 72 |
9 |
0.00185 |
| 40 |
8 |
0.00113 |
| 14 |
7 |
0.00101 |
| 8 |
6 |
0.00116 |
| 17 |
6 |
0.000646 |
| 9 |
5 |
0.000806 |
| 11 |
5 |
0.000705 |
| 27 |
5 |
0.00102 |
| 33 |
5 |
0.00102 |
| 41 |
5 |
0.000766 |
| 98 |
5 |
0.00111 |
| 5 |
4 |
0.000775 |
| 12 |
4 |
0.000526 |
| 16 |
4 |
0.000655 |
| 20 |
4 |
0.000573 |
| 22 |
4 |
0.000637 |
| 23 |
4 |
0.000676 |
| 29 |
4 |
0.000742 |
| 50 |
4 |
0.000766 |
| 57 |
4 |
0.000862 |
| 74 |
4 |
0.000792 |
| 96 |
4 |
0.000868 |
| 108 |
4 |
0.000904 |
| 114 |
4 |
0.000939 |
| 156 |
4 |
0.000897 |
| 175 |
4 |
0.000949 |
| 182 |
4 |
0.000907 |
| 213 |
4 |
0.000849 |
| 1 |
3 |
0.000425 |
| 3 |
3 |
0.000418 |
| 6 |
3 |
0.000409 |
| 10 |
3 |
0.000604 |
| 13 |
3 |
0.000594 |
| 15 |
3 |
0.000505 |
| 25 |
3 |
0.000554 |
| 28 |
3 |
0.000598 |
| 48 |
3 |
0.000442 |
| 52 |
3 |
0.000553 |
| 59 |
3 |
0.000639 |
| 61 |
3 |
0.000574 |
| 63 |
3 |
0.000458 |
| 65 |
3 |
0.000613 |
| 82 |
3 |
0.000653 |
| 95 |
3 |
0.000657 |
| 127 |
3 |
0.000624 |
| 153 |
3 |
0.000611 |
| 21 |
2 |
0.000339 |
| 24 |
2 |
0.000302 |
| 26 |
2 |
0.000354 |
| 30 |
2 |
0.000279 |
| 34 |
2 |
0.000337 |
| 36 |
2 |
0.000389 |
| 37 |
2 |
3e-04 |
| 42 |
2 |
0.000371 |
| 46 |
2 |
0.000388 |
| 49 |
2 |
0.000379 |
| 60 |
2 |
0.000428 |
| 69 |
2 |
0.000429 |
| 70 |
2 |
0.000447 |
| 77 |
2 |
0.000411 |
| 79 |
2 |
0.000334 |
| 81 |
2 |
0.000343 |
| 84 |
2 |
0.000388 |
| 89 |
2 |
0.000407 |
| 90 |
2 |
0.000334 |
| 92 |
2 |
0.000472 |
| 109 |
2 |
0.000463 |
| 110 |
2 |
0.000445 |
| 122 |
2 |
0.000414 |
| 125 |
2 |
0.000457 |
| 132 |
2 |
0.000457 |
| 139 |
2 |
0.000424 |
| 143 |
2 |
0.000416 |
| 154 |
2 |
0.000479 |
| 161 |
2 |
0.000406 |
| 163 |
2 |
0.000408 |
| 166 |
2 |
0.000465 |
| 187 |
2 |
0.00047 |
| 205 |
2 |
0.000469 |
| 211 |
2 |
0.000483 |
| 255 |
2 |
0.00049 |
| 6450 |
2 |
0.000549 |
| 32900 |
2 |
0.000549 |
| 66600 |
2 |
0.000549 |
| 259000 |
2 |
0.000549 |
| 2 |
1 |
0.00011 |
| 7 |
1 |
0.000124 |
| 18 |
1 |
0.000185 |
| 32 |
1 |
0.000197 |
| 44 |
1 |
0.000152 |
| 45 |
1 |
0.000161 |
| 47 |
1 |
0.000165 |
| 56 |
1 |
0.000259 |
| 62 |
1 |
0.00016 |
| 68 |
1 |
0.000198 |
| 71 |
1 |
0.000171 |
| 75 |
1 |
0.000153 |
| 78 |
1 |
0.000164 |
| 83 |
1 |
0.000209 |
| 85 |
1 |
0.000203 |
| 86 |
1 |
0.000166 |
| 87 |
1 |
0.000203 |
| 88 |
1 |
0.00017 |
| 91 |
1 |
0.000233 |
| 93 |
1 |
0.000228 |
| 97 |
1 |
0.000194 |
| 99 |
1 |
0.000233 |
| 100 |
1 |
0.00022 |
| 103 |
1 |
0.000179 |
| 104 |
1 |
0.000225 |
| 105 |
1 |
0.000198 |
| 106 |
1 |
2e-04 |
| 111 |
1 |
0.000163 |
| 116 |
1 |
0.000177 |
| 123 |
1 |
0.000215 |
| 126 |
1 |
0.000211 |
| 130 |
1 |
0.000177 |
| 131 |
1 |
0.000256 |
| 134 |
1 |
0.000206 |
| 138 |
1 |
0.00023 |
| 144 |
1 |
0.000242 |
| 149 |
1 |
0.000232 |
| 152 |
1 |
0.000187 |
| 155 |
1 |
0.000229 |
| 157 |
1 |
0.000227 |
| 159 |
1 |
0.000226 |
| 162 |
1 |
0.000209 |
| 164 |
1 |
0.00026 |
| 171 |
1 |
0.000224 |
| 179 |
1 |
0.000236 |
| 180 |
1 |
0.000247 |
| 181 |
1 |
0.000229 |
| 189 |
1 |
0.000218 |
| 193 |
1 |
0.000245 |
| 194 |
1 |
0.000262 |
| 195 |
1 |
0.00025 |
| 200 |
1 |
0.000224 |
| 206 |
1 |
0.000245 |
| 209 |
1 |
0.00026 |
| 217 |
1 |
0.000241 |
| 222 |
1 |
0.00025 |
| 224 |
1 |
0.000268 |
| 238 |
1 |
0.000255 |
| 245 |
1 |
0.000236 |
| 260 |
1 |
0.000261 |
| 273 |
1 |
0.000246 |
| 280 |
1 |
0.000261 |
| 282 |
1 |
0.000264 |
| 361 |
1 |
0.000268 |
| 390 |
1 |
0.000271 |
| 449 |
1 |
0.000271 |
| 453 |
1 |
0.000269 |
| 481 |
1 |
0.00027 |
| 626 |
1 |
0.000273 |
| 865 |
1 |
0.000273 |
| 1160 |
1 |
0.000274 |
| 1550 |
1 |
0.000273 |
| 1610 |
1 |
0.000274 |
| 1870 |
1 |
0.000274 |
| 1870 |
1 |
0.000274 |
| 2500 |
1 |
0.000274 |
| 3640 |
1 |
0.000274 |
| 5640 |
1 |
0.000274 |
| 6040 |
1 |
0.000274 |
| 7850 |
1 |
0.000274 |
| 8100 |
1 |
0.000274 |
| 8870 |
1 |
0.000274 |
| 10800 |
1 |
0.000274 |
| 11300 |
1 |
0.000275 |
| 12300 |
1 |
0.000274 |
| 12900 |
1 |
0.000275 |
| 13000 |
1 |
0.000274 |
| 13200 |
1 |
0.000274 |
| 14900 |
1 |
0.000274 |
| 15800 |
1 |
0.000274 |
| 17800 |
1 |
0.000274 |
| 20500 |
1 |
0.000274 |
| 23900 |
1 |
0.000274 |
| 24300 |
1 |
0.000274 |
| 25800 |
1 |
0.000274 |
| 32600 |
1 |
0.000275 |
| 45900 |
1 |
0.000275 |
| 52300 |
1 |
0.000274 |
| 58800 |
1 |
0.000275 |
| 76000 |
1 |
0.000274 |
| 77000 |
1 |
0.000275 |
| 79900 |
1 |
0.000275 |
| 83000 |
1 |
0.000274 |
| 84700 |
1 |
0.000274 |
| 86100 |
1 |
0.000275 |
| 94500 |
1 |
0.000275 |
| 115000 |
1 |
0.000275 |
| 118000 |
1 |
0.000275 |
| 118000 |
1 |
0.000275 |
| 123000 |
1 |
0.000275 |
| 138000 |
1 |
0.000275 |
| 139000 |
1 |
0.000275 |
| 177000 |
1 |
0.000275 |
| 185000 |
1 |
0.000275 |
| 188000 |
1 |
0.000275 |
| 192000 |
1 |
0.000275 |
| 198000 |
1 |
0.000275 |
| 202000 |
1 |
0.000275 |
| 205000 |
1 |
0.000275 |
| 228000 |
1 |
0.000275 |
| 231000 |
1 |
0.000275 |
| 232000 |
1 |
0.000275 |
| 234000 |
1 |
0.000275 |
| 244000 |
1 |
0.000275 |
| 253000 |
1 |
0.000275 |
| 266000 |
1 |
0.000275 |
| 295000 |
1 |
0.000275 |
| 311000 |
1 |
0.000275 |
| 326000 |
1 |
0.000275 |
| 337000 |
1 |
0.000275 |
|
CL55 ----> CL35
No. of shared pairs: :2535
CL55 ----> CL53
No. of shared pairs: :2282
CL55 ----> CL67
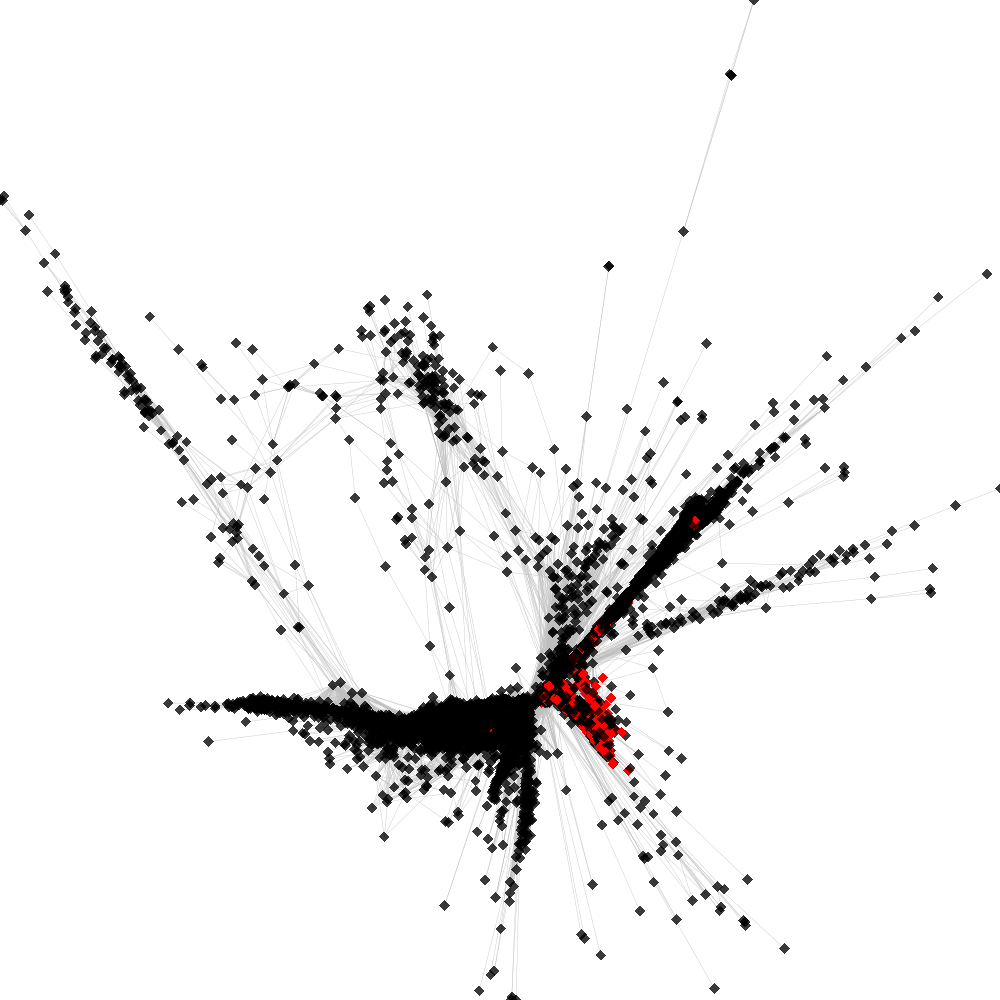

No. of shared pairs: :878
CL55 ----> CL190

No. of shared pairs: :619
CL55 ----> CL43
No. of shared pairs: :123
CL55 ----> CL117
No. of shared pairs: :72
CL55 ----> CL524
No. of shared pairs: :22
CL55 ----> CL31

No. of shared pairs: :14
CL55 ----> CL4

No. of shared pairs: :10
CL55 ----> CL102

No. of shared pairs: :10

