| |
kmer |
variant |
total_score |
monomer_length |
score_bn |
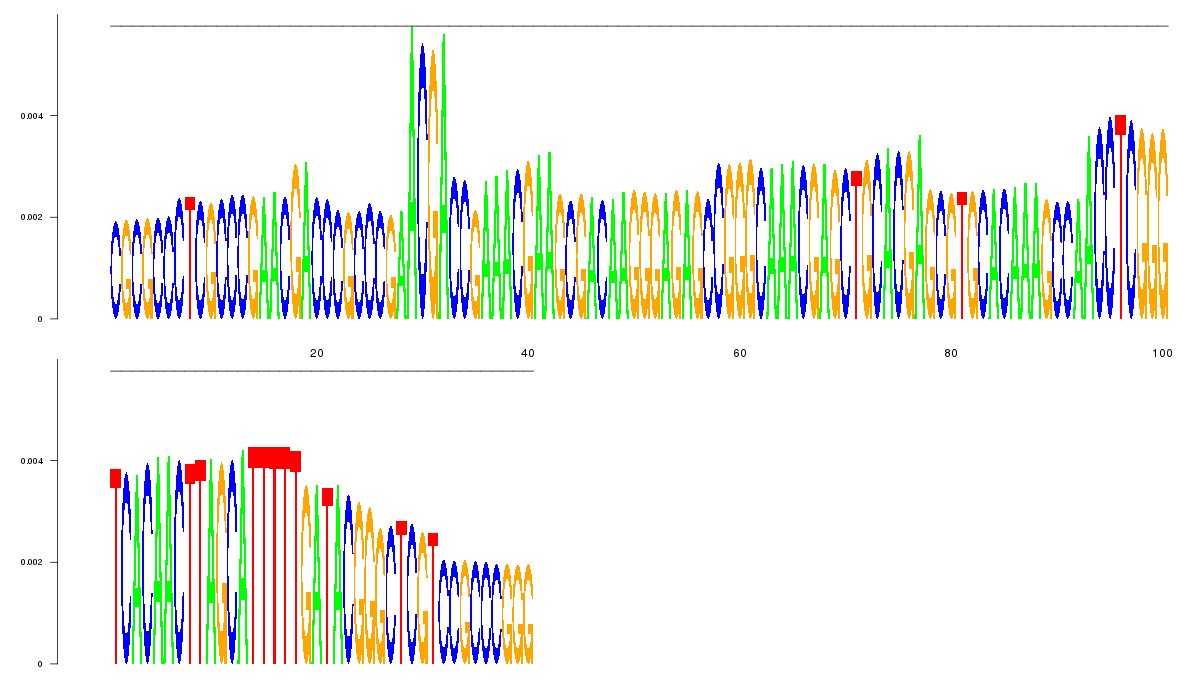
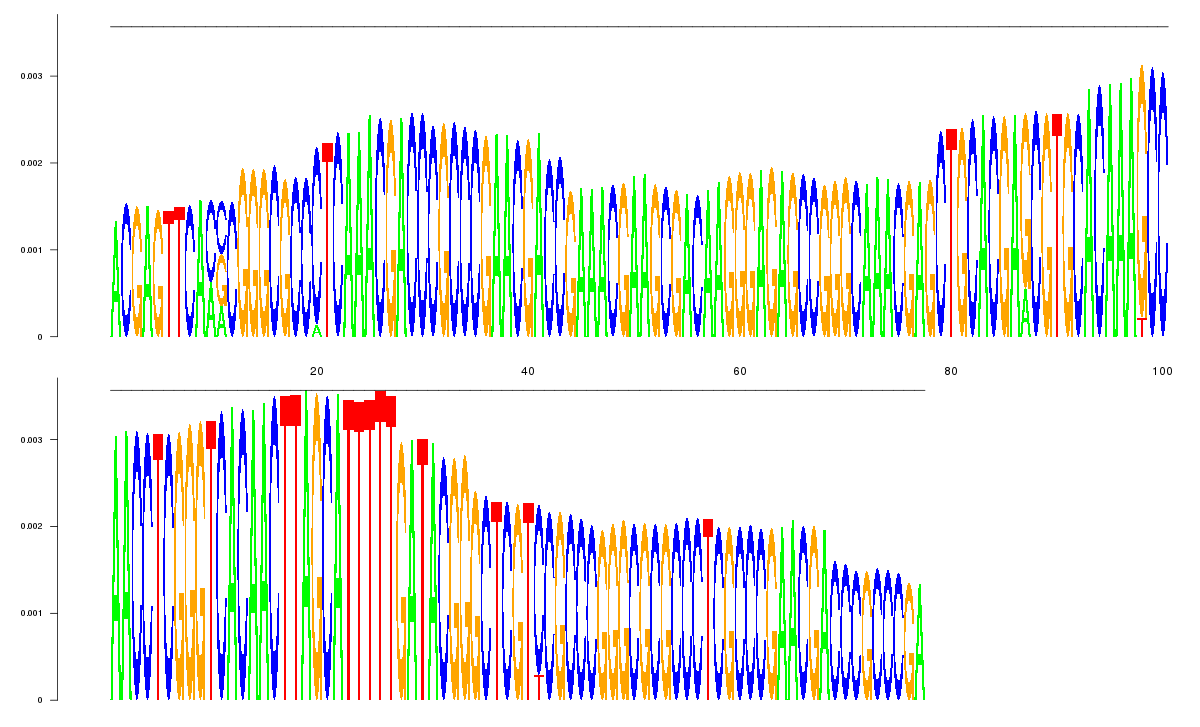
consensus |
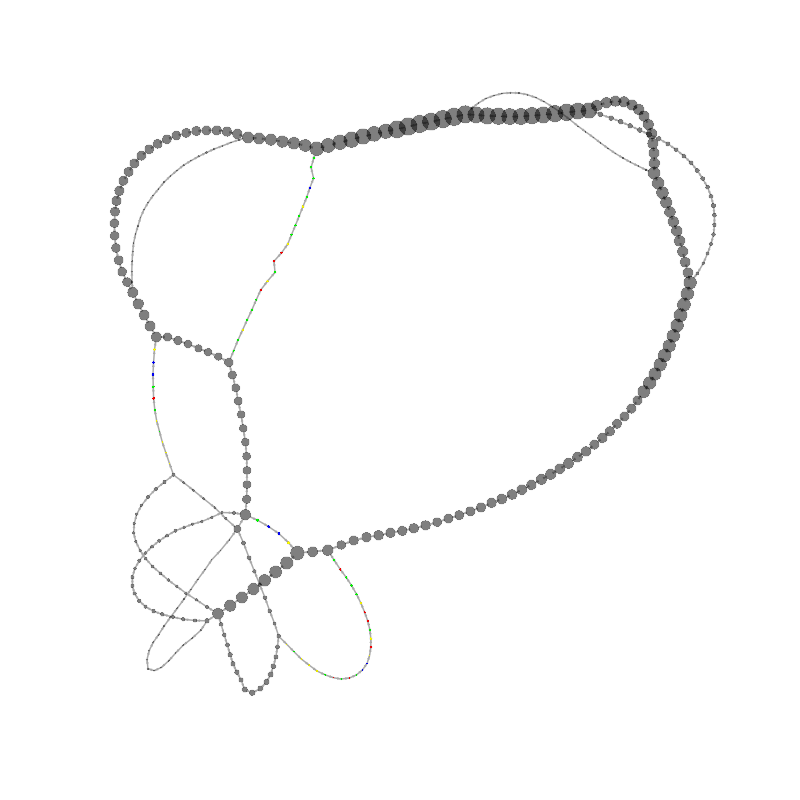
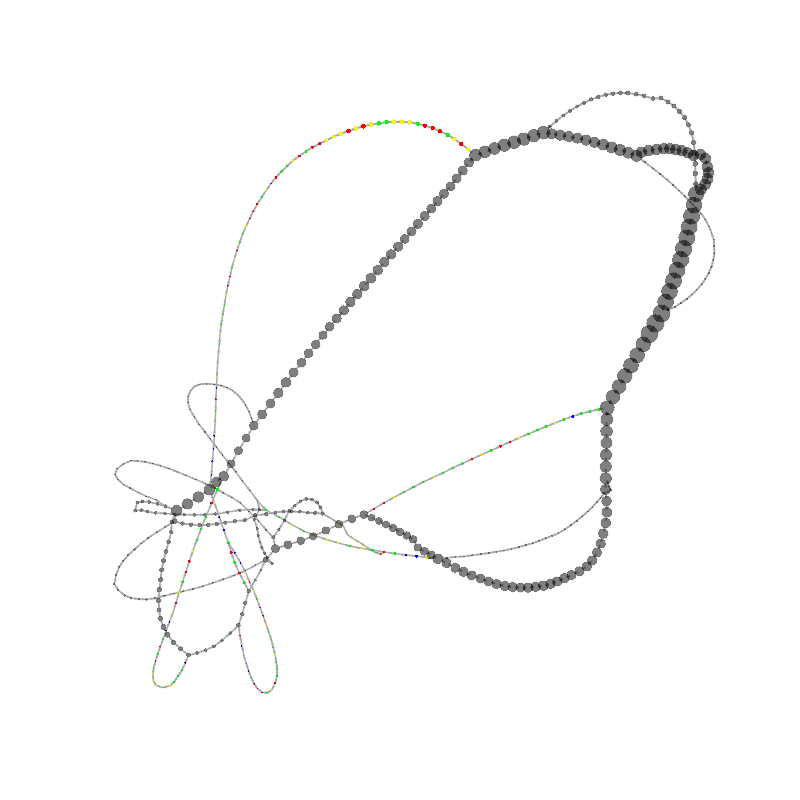
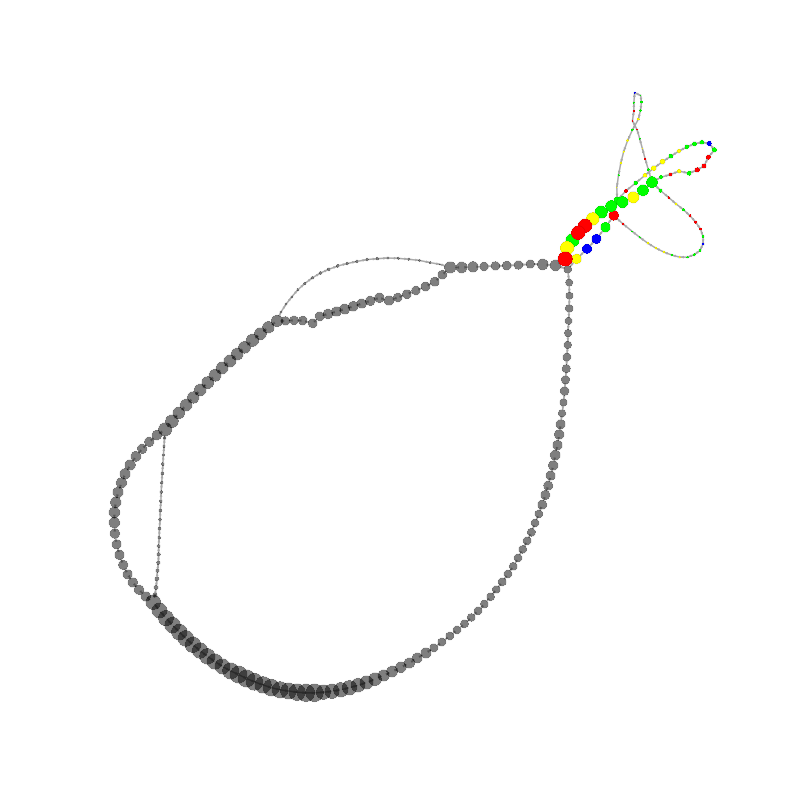
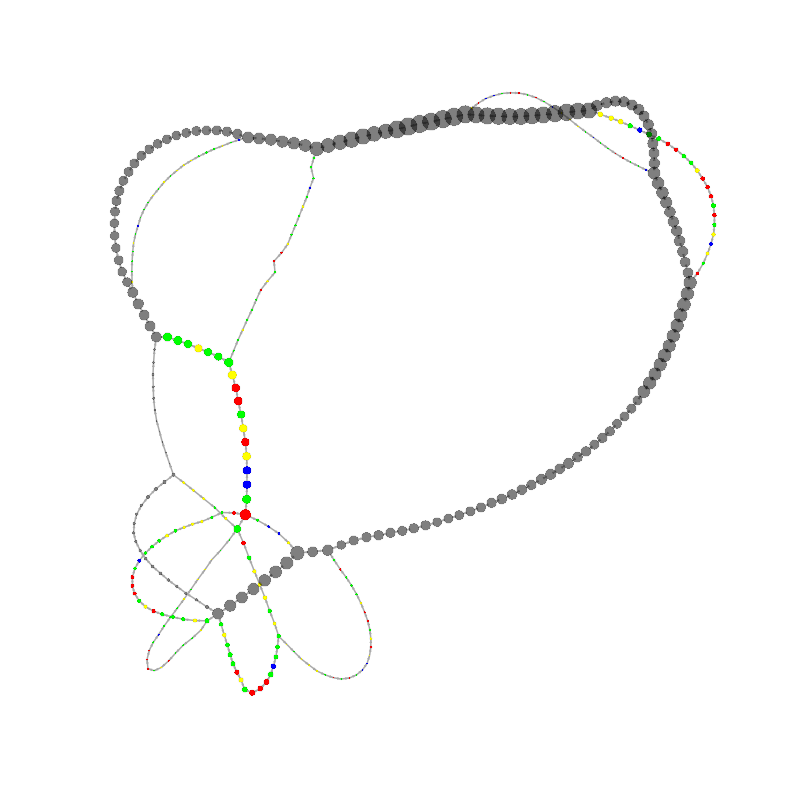
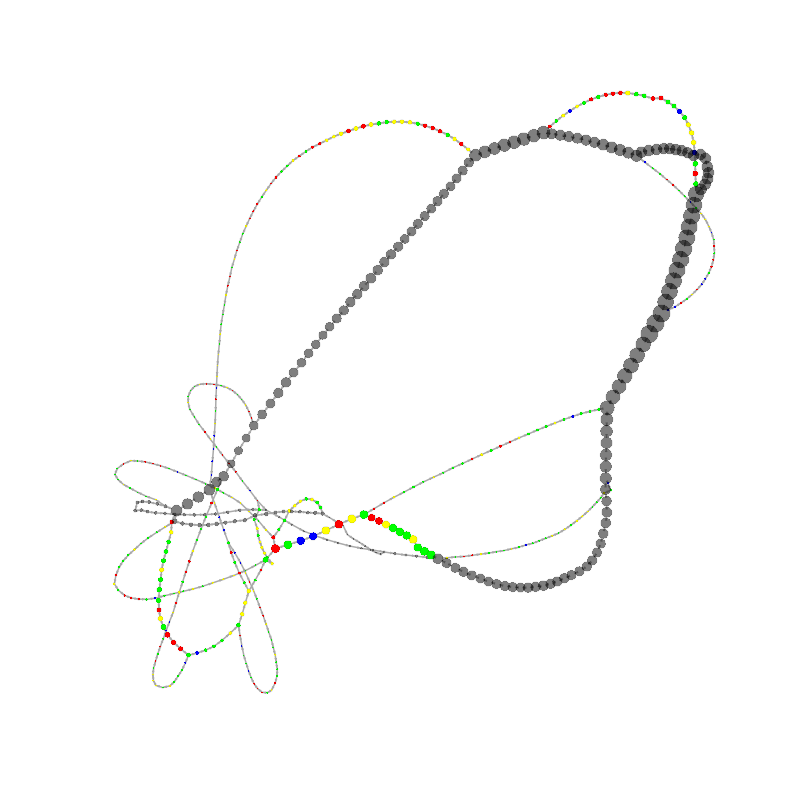
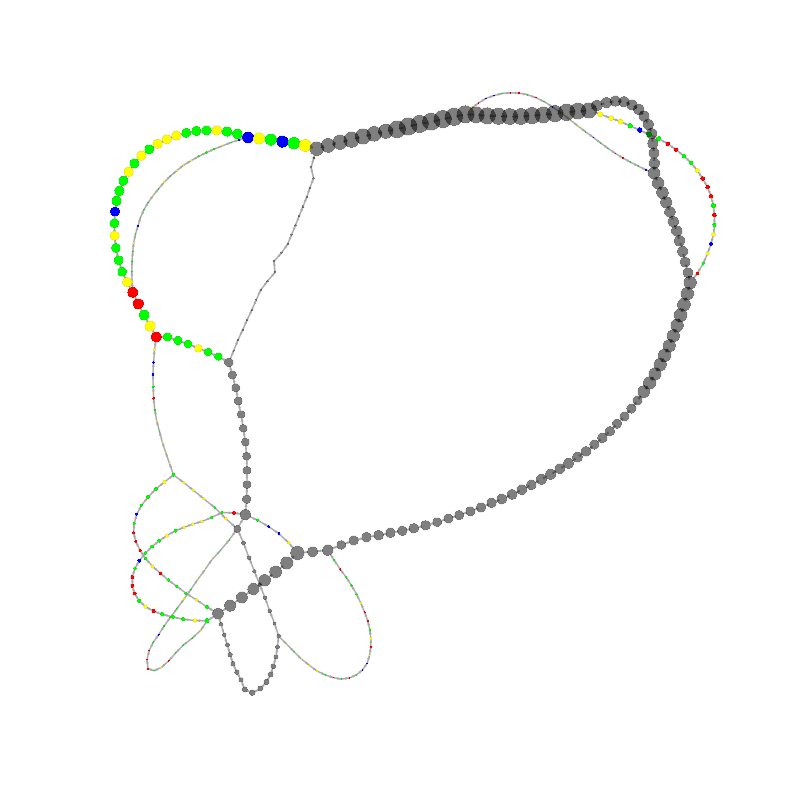
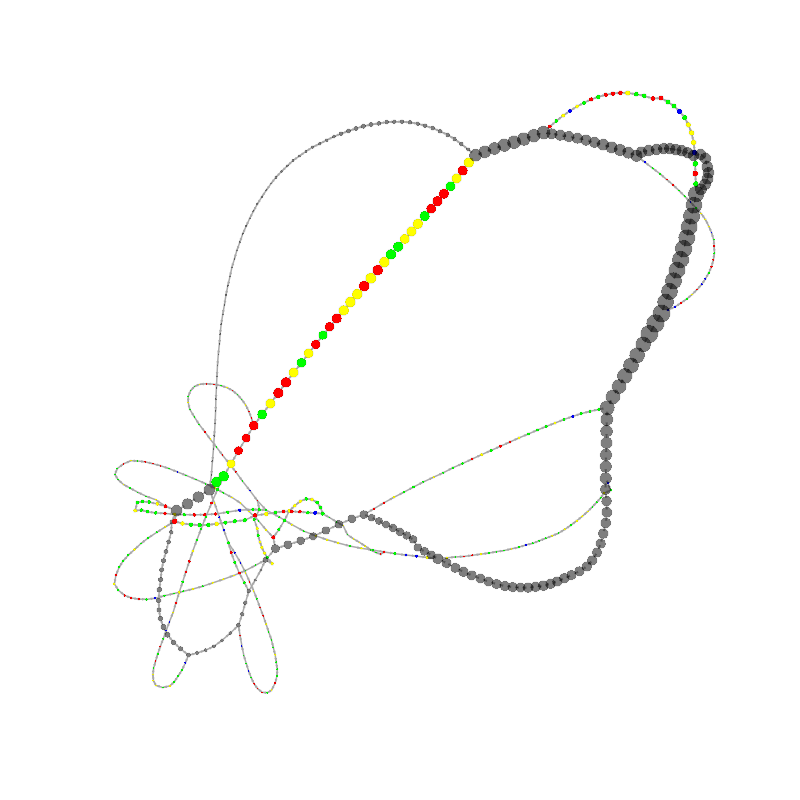
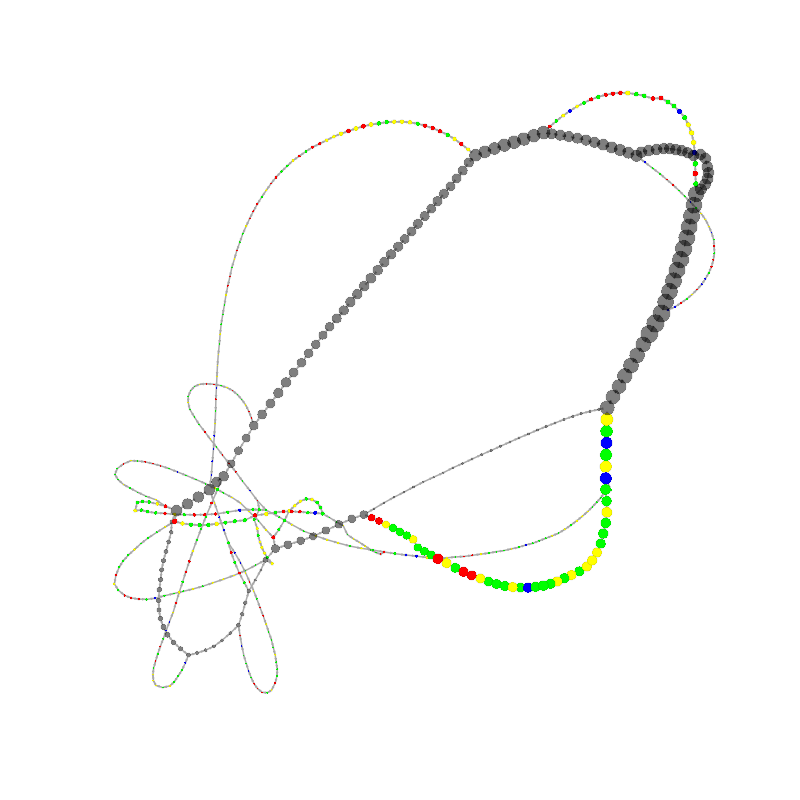
graph_image |
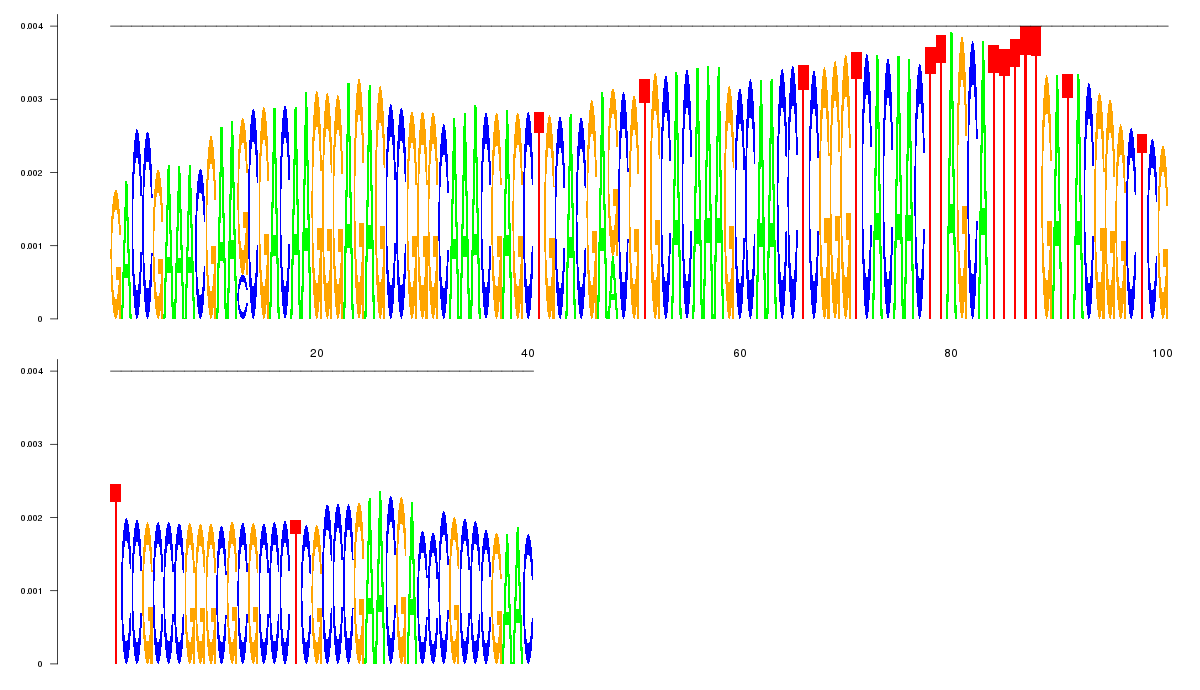
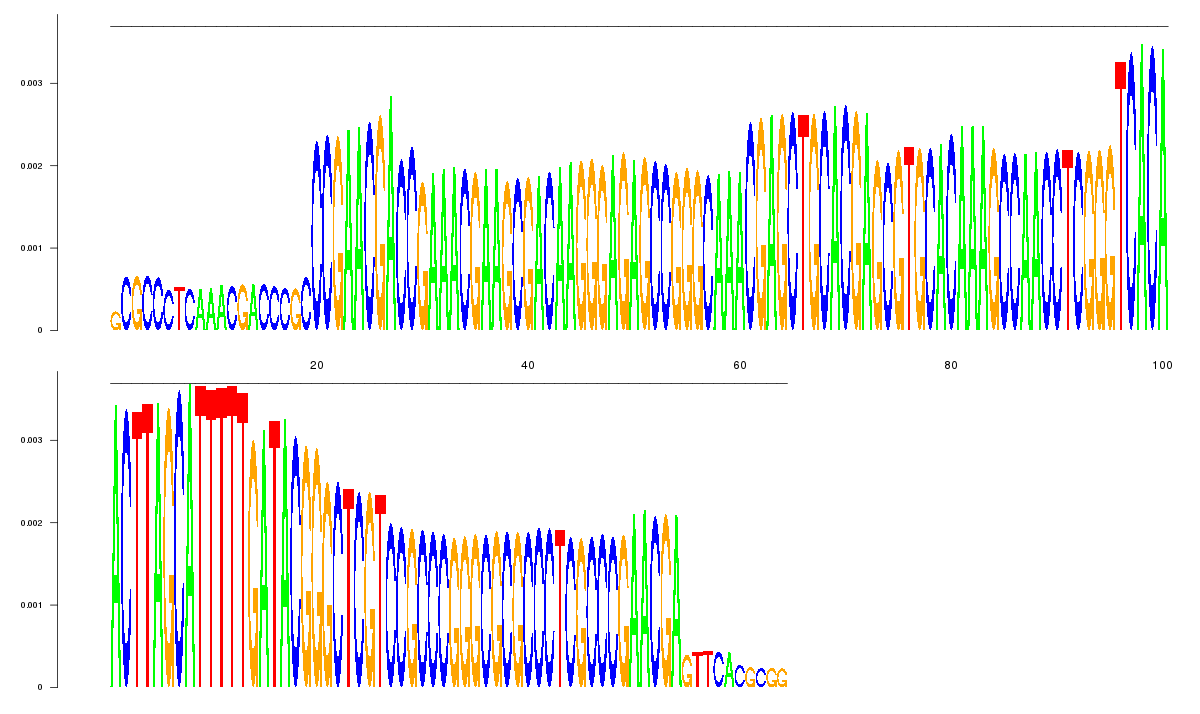
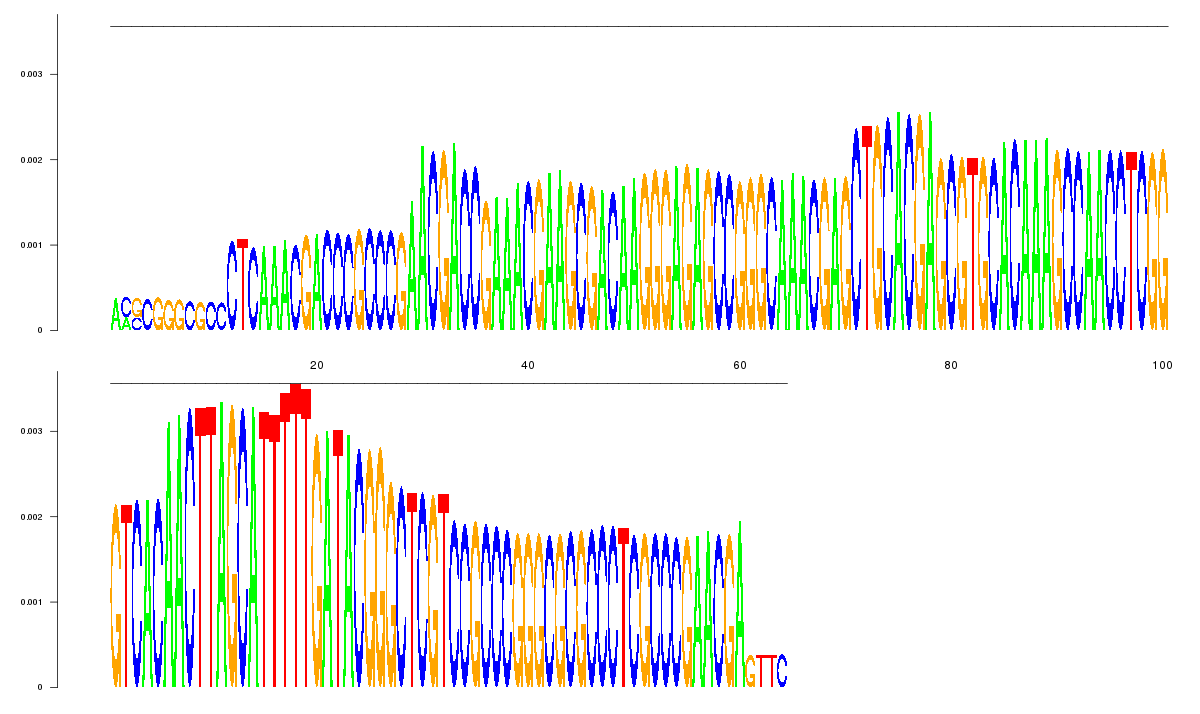
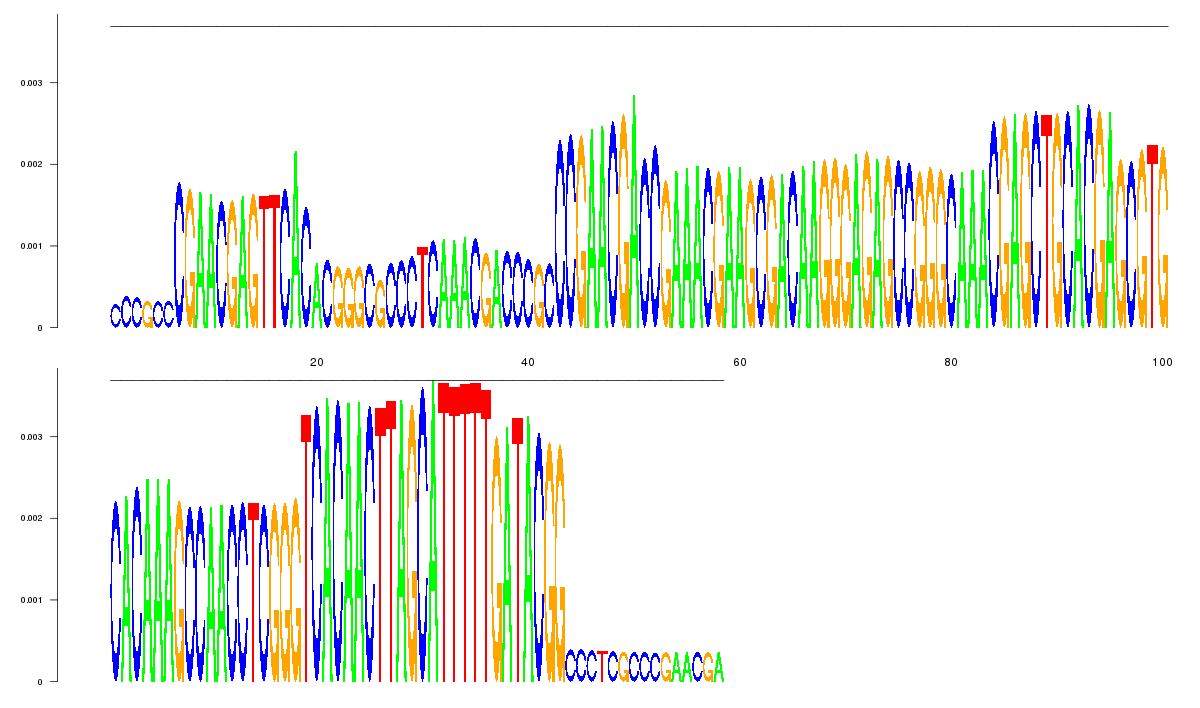
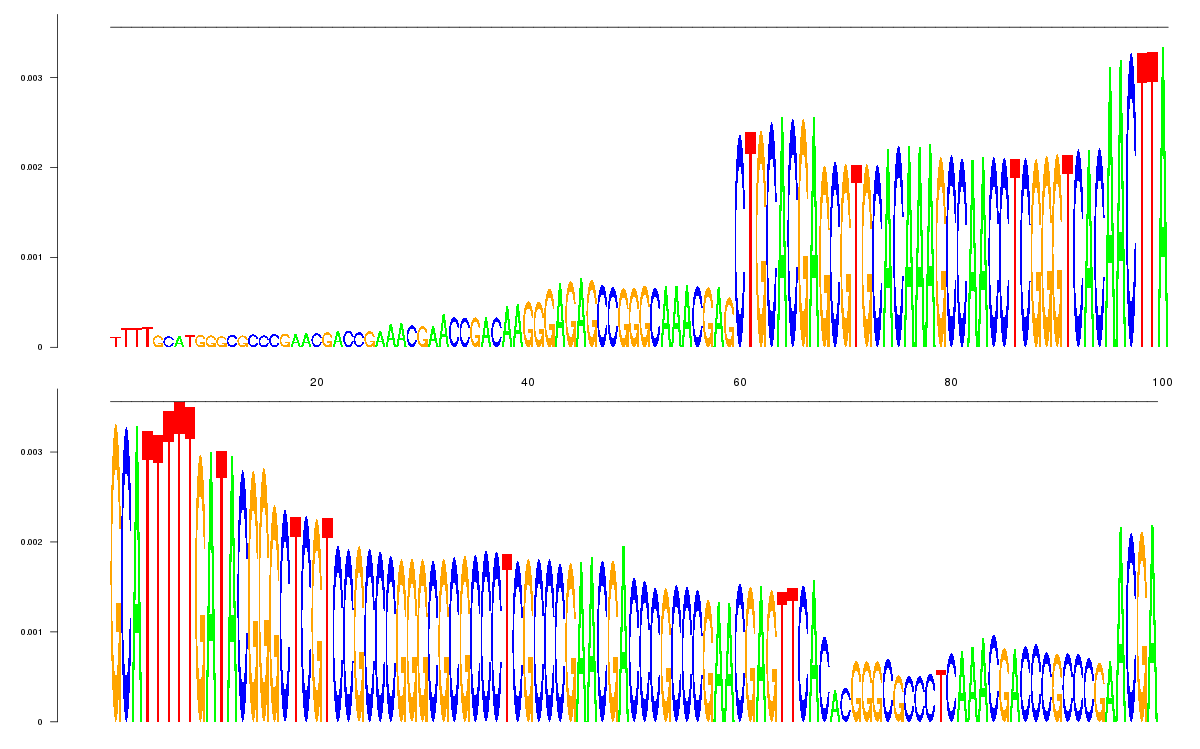
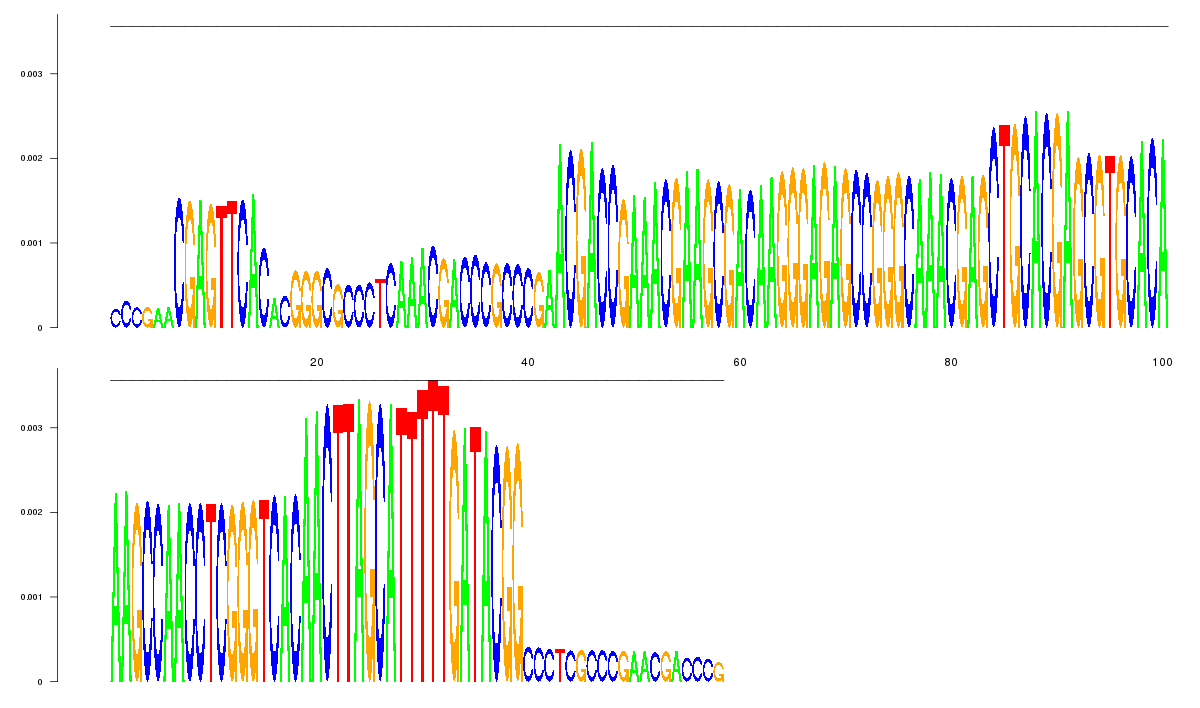
logo_image |
| 1
|
11
|
3
|
0.464
|
127
|
0.00246
|
CGCCCTCGCCCGAACGACCGAAACGAAGCGACAAGGGAGAGCCGGGCAAACGAGCTGCACGAGCGTGCACAAAGCCAACC
TCGGGTCACAACTTAGCATTTTTGATACGGGCTCGTCCGCCCGGGCG
|
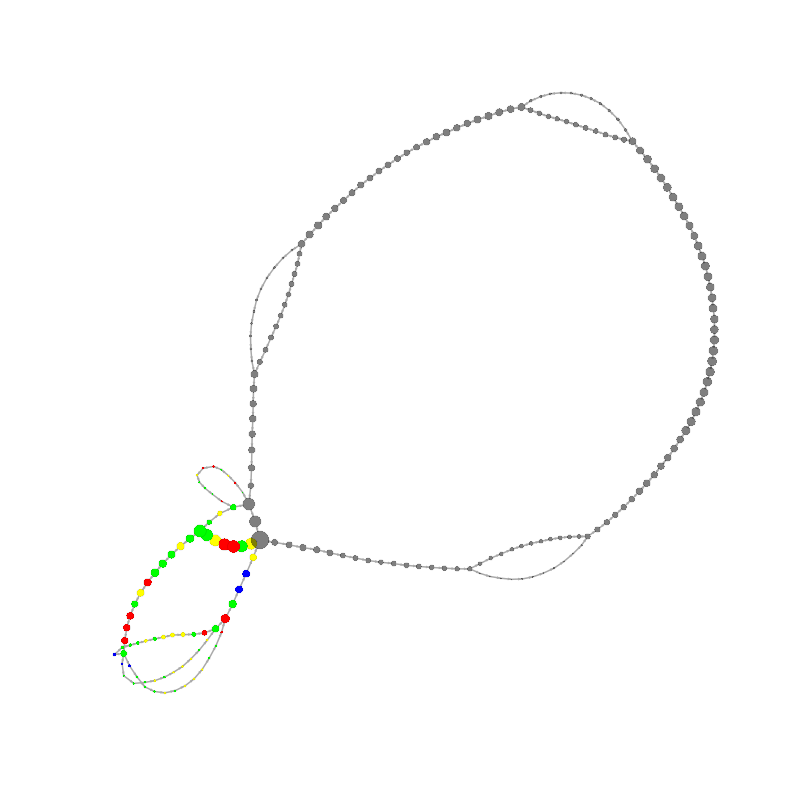
|
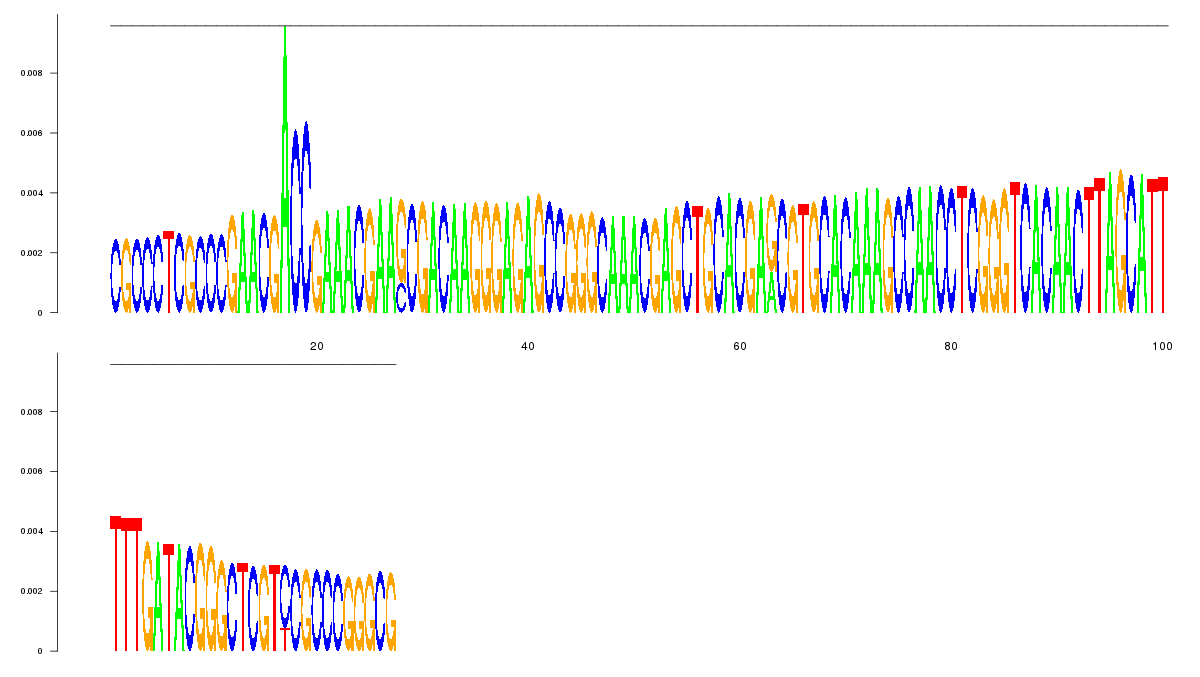
|
| 2
|
23
|
5
|
0.436
|
177
|
0.00153
|
GCCCGAACGAGTTCACACGGGCGCCCTCAAACGACCCGCCCGAACGACCGAAACGAAGCGACAAGGGAGAGCCGGGCAAA
CGAGCTGCACGAGCGTGCACAAAGCCAACCTCGGGTCACAACTTAGCATTTTTGATACGGGCTCGTCCGCCCGGGCGCGC
CCTCGCCCGAACGACCC
|

|
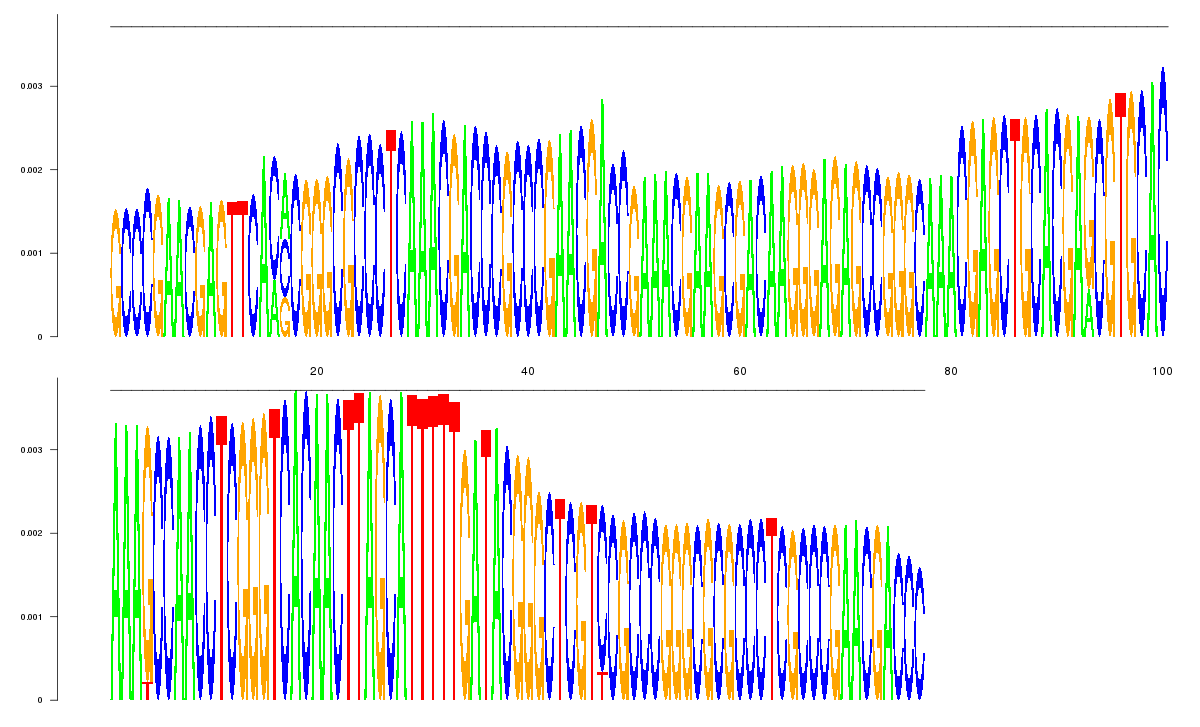
|
| 3
|
15
|
2
|
0.413
|
140
|
0.00192
|
CGCGCCCTCGCCCGAACGACCCGCCCGAACGACCGAAACGAAGCGACAAGGGAGAGCCGGGCAAACGAGCTGCACGAGCG
TGCACAAAGCCAACCTCGGGTCACAACTTAGCATTTTTGATACGGGCTCGTCCGCCCGGG
|
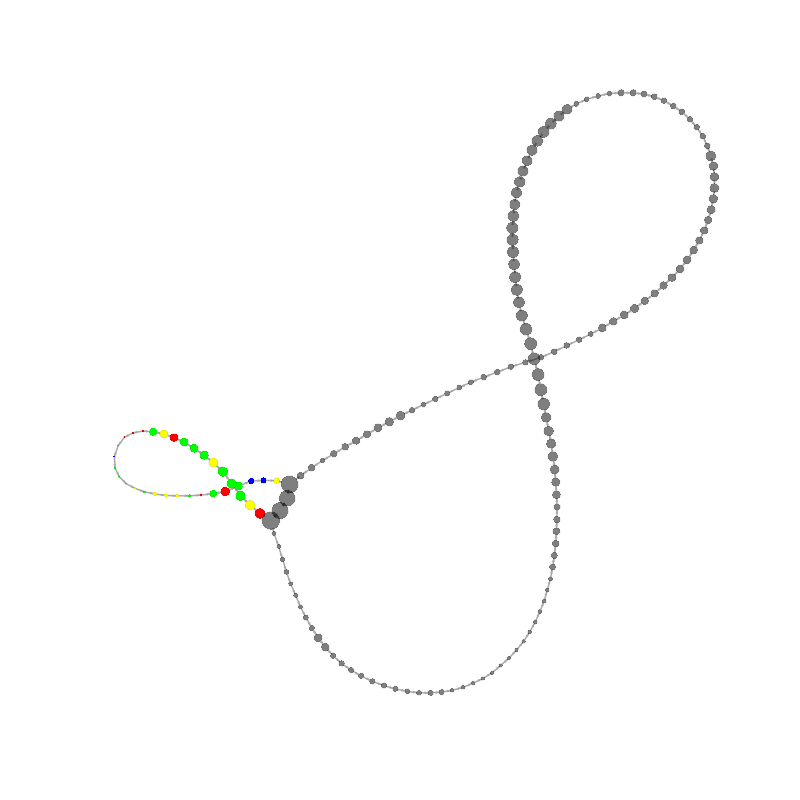
|
|
| 4
|
27
|
5
|
0.405
|
177
|
0.00133
|
ACGAGTTCACCCGGGCGCCCTCAAACGACCCGCCCGAACGACCGAAACGAAGCGACAAGGGAGAGCCGGGCAAACGAGCT
GCACGAGCGTGCACAAAGCCAACCTCGGGTCACAACTTAGCATTTTTGATACGGGCTCGTCCGCCCGGGCGCGCCCTCGC
CCGAACGACCCGCCCGA
|
|
|
| 5
|
19
|
2
|
0.388
|
140
|
0.00176
|
GACCGAAACGAAGCGACAAGGGAGAGCCGGGCAAACGAGCTGCACGAGCGTGCACAAAGCCAACCTCGGGTCACAACTTA
GCATTTTTGATACGGGCTCGTCCGCCCGGGCGCGCCCTCGCCCGAACGACCCGCCCGAAC
|
|
|
| 6
|
23
|
4
|
0.335
|
164
|
0.00023
|
GCGCCCTCAAACGACCCGCCCGAACGACCGAAACGAAGCGACAAGGGAGAGCCGGGCAAACGAGCTGCACGAGCGTGCAC
AAAGCCAACCTCGGGTCACAACTTAGCATTTTTGATACGGGCTCGTCCGCCCGGGCGCGCCCTCGCCCGAACGAGTTCAC
GCGG
|
|
|
| 7
|
27
|
4
|
0.310
|
164
|
0.00033
|
ACGCGGGCGCCCTCAAACGACCCGCCCGAACGACCGAAACGAAGCGACAAGGGAGAGCCGGGCAAACGAGCTGCACGAGC
GTGCACAAAGCCAACCTCGGGTCACAACTTAGCATTTTTGATACGGGCTCGTCCGCCCGGGCGCGCCCTCGCCCGAACGA
GTTC
|
|
|
| 8
|
23
|
3
|
0.302
|
158
|
0.00029
|
CCCGCCCGAACGAGTTCACACGGGCGCCCTCAAACGACCCGCCCGAACGACCGAAACGAAGCGACAAGGGAGAGCCGGGC
AAACGAGCTGCACGAGCGTGCACAAAGCCAACCTCGGGTCACAACTTAGCATTTTTGATACGGCCCTCGCCCGAACGA
|
|
|
| 9
|
27
|
6
|
0.286
|
199
|
0.00011
|
TTTTGCATGGGCGCCCGAACGACCGAAACGAACCGACAAGGGAGAGCCGGGCAAACGAGCTGCACGAGCGTGCACAAAGC
CAACCTCGGGTCACAACTTAGCATTTTTGATACGGGCTCGTCCGCCCGGGCGCGCCCTCGCCCGAACGACCCGCCCGAAC
GAGTTCACACGGGCGCCCTCAAACGACCCGCCCGAACGA
|
|
|
| 10
|
27
|
3
|
0.259
|
158
|
0.00022
|
CCCGAACGAGTTCACACGGGCGCCCTCAAACGACCCGCCCGAACGACCGAAACGAAGCGACAAGGGAGAGCCGGGCAAAC
GAGCTGCACGAGCGTGCACAAAGCCAACCTCGGGTCACAACTTAGCATTTTTGATACGGCCCTCGCCCGAACGACCCG
|
|
|
| 11
|
11
|
2
|
0.163
|
37
|
0.00302
|
CAAACGACCCGCCCGAACGAGTTCACACGGGCGCCCT
|
|
|
| 12
|
15
|
1
|
0.100
|
37
|
0.00126
|
GCCCTCAAACGACCCGCCCGAACGAGTTCACACGGGC
|
|
|
| 13
|
19
|
1
|
0.096
|
37
|
0.00199
|
TCACACGGGCGCCCTCAAACGACCCGCCCGAACGAGT
|
|
|
| 14
|
11
|
1
|
0.082
|
13
|
0.00306
|
CGCCCGAACGACC
|
|
|
| 15
|
23
|
2
|
0.046
|
50
|
0.00019
|
GCGCCCTCAAACGACCCGCCCGAACGACCCACCCGAACGAGTTCACACGG
|
|
|
| 16
|
23
|
1
|
0.046
|
37
|
0.00045
|
TCACACGGGCGCCCTCAAACGACCCGCCCGAACGAGT
|
|
|
| 17
|
27
|
1
|
0.034
|
37
|
0.00037
|
CGGGCGCCCTCAAACGACCCGCCCGAACGAGTTCACA
|
|
|
| 18
|
27
|
2
|
0.032
|
50
|
0.00012
|
TTCACACGGGCGCCCTCAAACGACCCGCCCGAACGACCCACCCGAACGAG
|
|
|